VERO Researcher Studies Common Pathogen To Improve Diagnosis, Treatment Of Bovine Respiratory Disease
Story by Aubrey Bloom, VMBS Communications

One of the primary research areas for the Texas A&M School of Veterinary Medicine & Biomedical Sciences’ (VMBS) Veterinary Research, Education, and Outreach (VERO) program is trying to solve the complex problem of Bovine Respiratory Disease (BRD).
Though research has been ongoing for decades, BRD still accounts for more than half of all feedlot deaths and remains one of the single largest economic impacts to the beef cattle industry, with some estimates that BRD costs the industry more than a billion dollars annually.
In an effort to better understand this complex problem, VMBS assistant professor Dr. Robert Valeris-Chacin wants to go back to the basics of understanding how exactly the disease manifests. He was recently awarded a $100,000 grant through Texas A&M AgriLife Research from the USDA-NIFA Animal Health and Disease Research Capacity Program to develop a new diagnostic tool for identifying specific strains of one of the common BRD-associated pathogens, Mycoplasma bovis.
One of the primary reasons BRD is a difficult problem to solve is that it isn’t caused by any one specific virus or bacteria but, instead, a number of different combinations of pathogens; the exact roles those pathogens play and how they interact with one another to cause BRD is still mostly a mystery.
That’s why Valeris-Chacin is trying to lay the groundwork for a more thorough understanding of those roles, and he’s starting specifically with M. bovis, a bacterium commonly found and associated with BRD.
According to Valeris-Chacin, M. bovis research lags behind some of the other BRD-associated pathogens for a couple of reasons.
One is that mycoplasmas are relatively hard to culture in a lab. Mycoplasmas require a significant amount of nutrients, and other bacteria will often outcompete mycoplasmas for those nutrients. As a result, many scientists have chosen to work on the pathogens that are easier to culture.
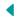
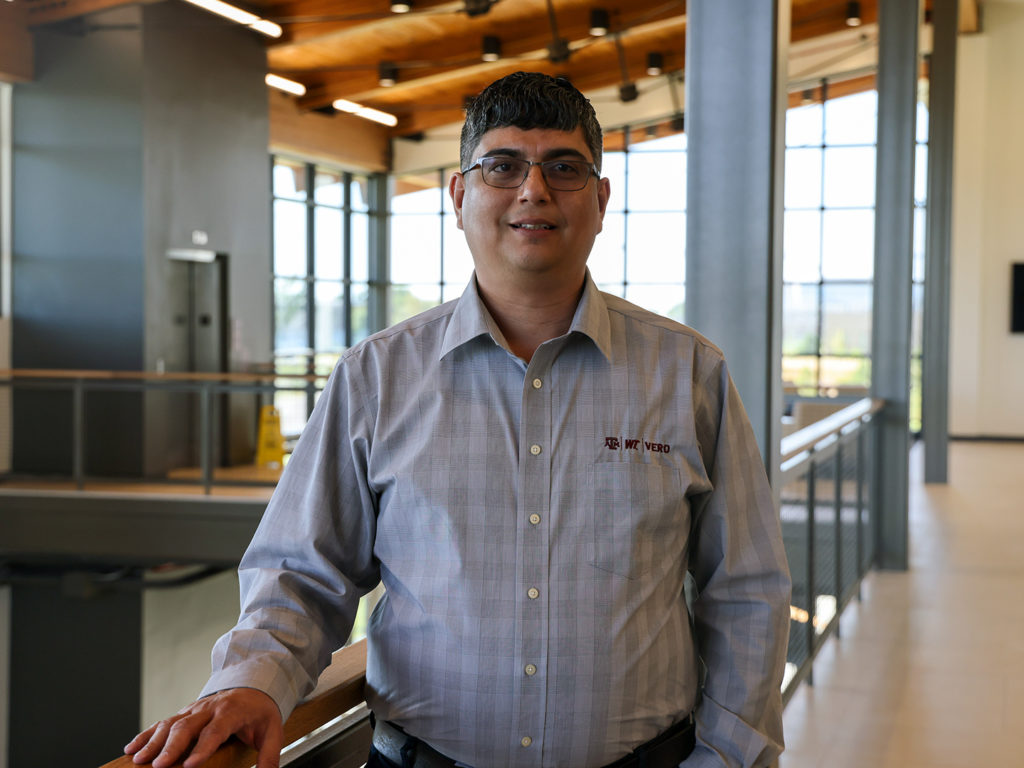
Another reason is that when infected animals are sampled, there is usually a very small amount of M. bovis in the sample. With current techniques, this means that it’s practically impossible to gather genetic information of M. bovis beyond whether the bacterium is present in the sample.
“Are there different strains of M. bovis present? Do they have differences in how they cause damage or how pathogenic they are? Those are questions that we just don’t have the answers to right now,” he said. “Current tests can tell us if the M. bovis is present but not a specific strain or if there are multiple strains.”
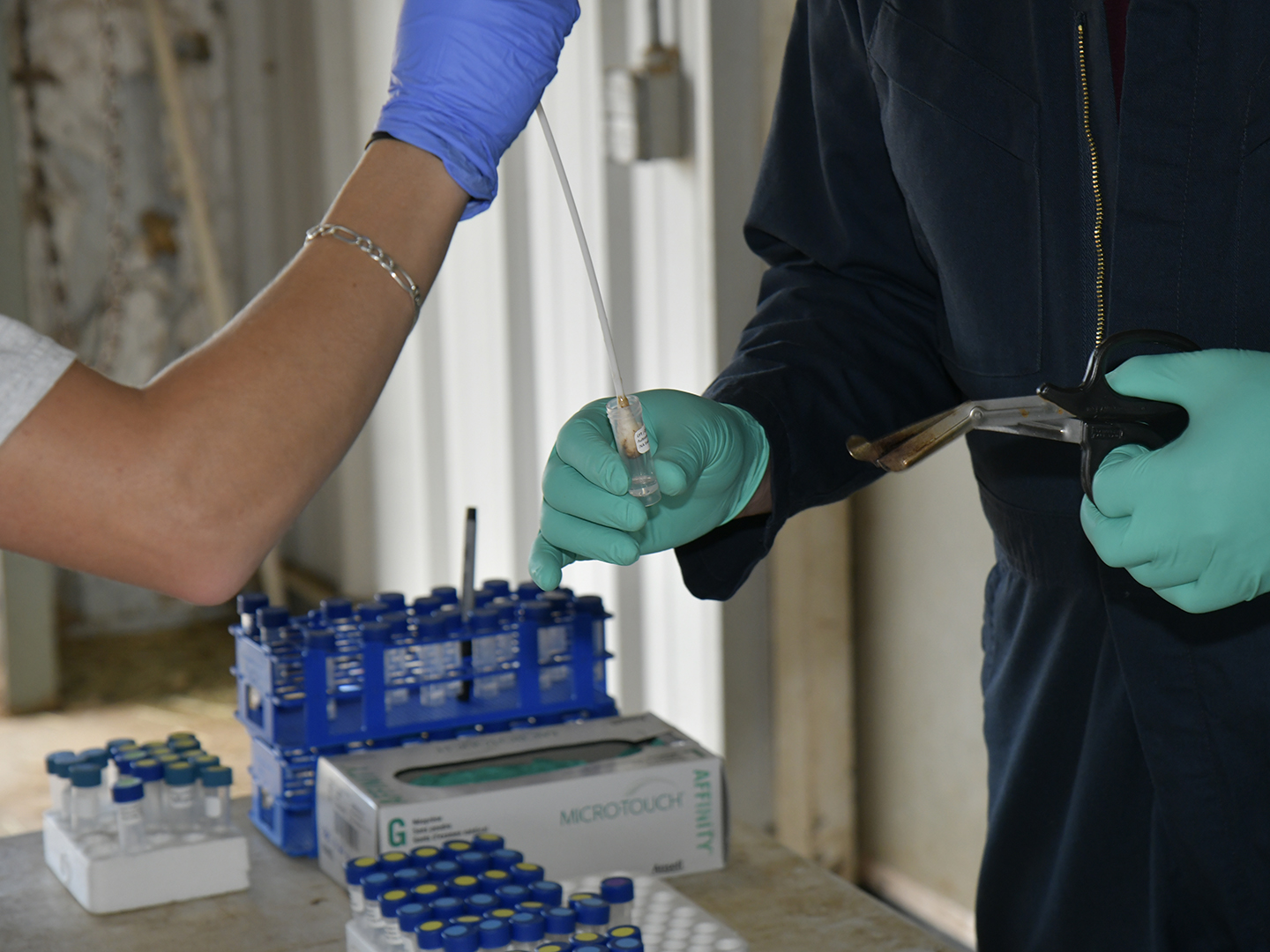
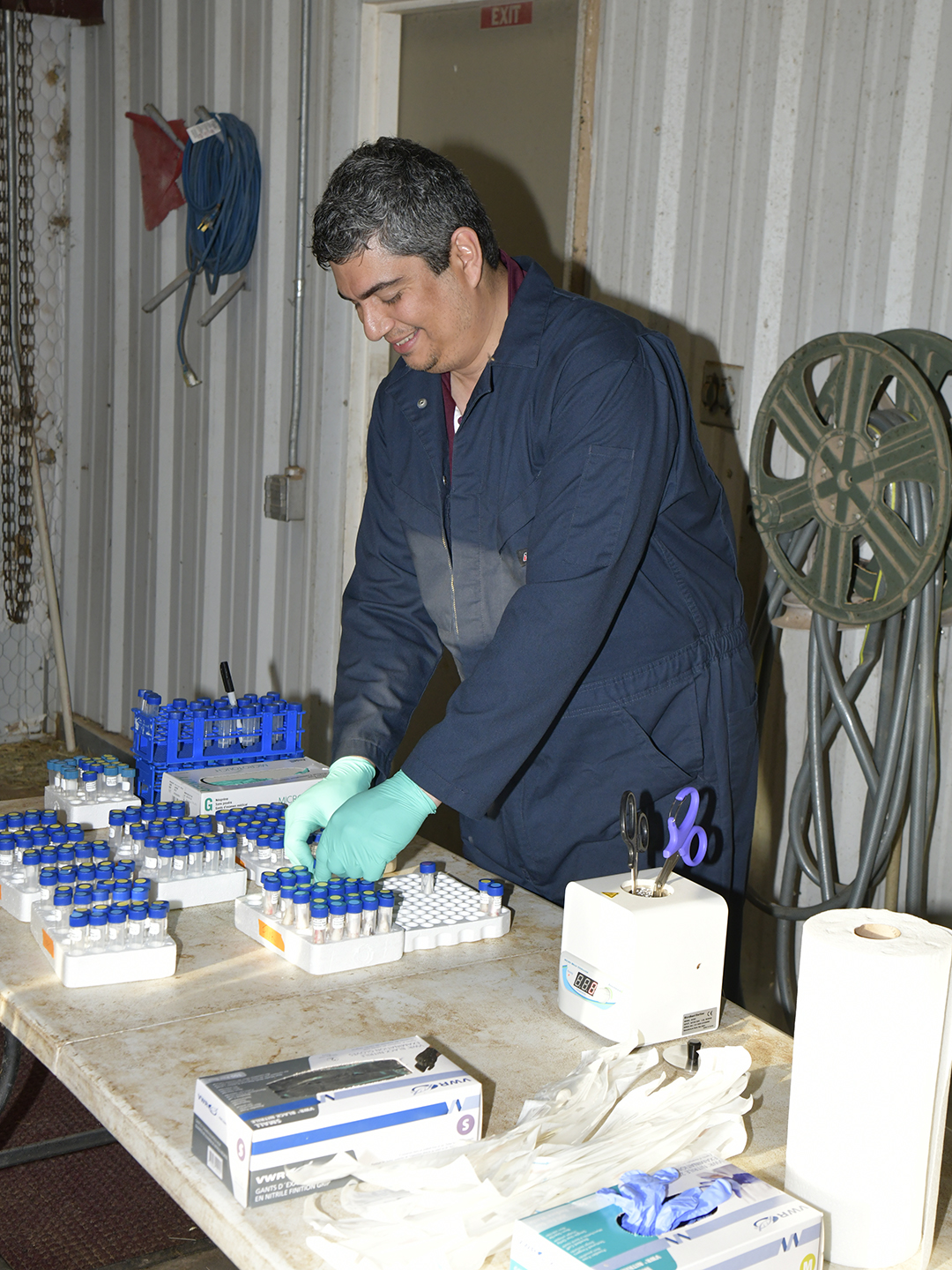
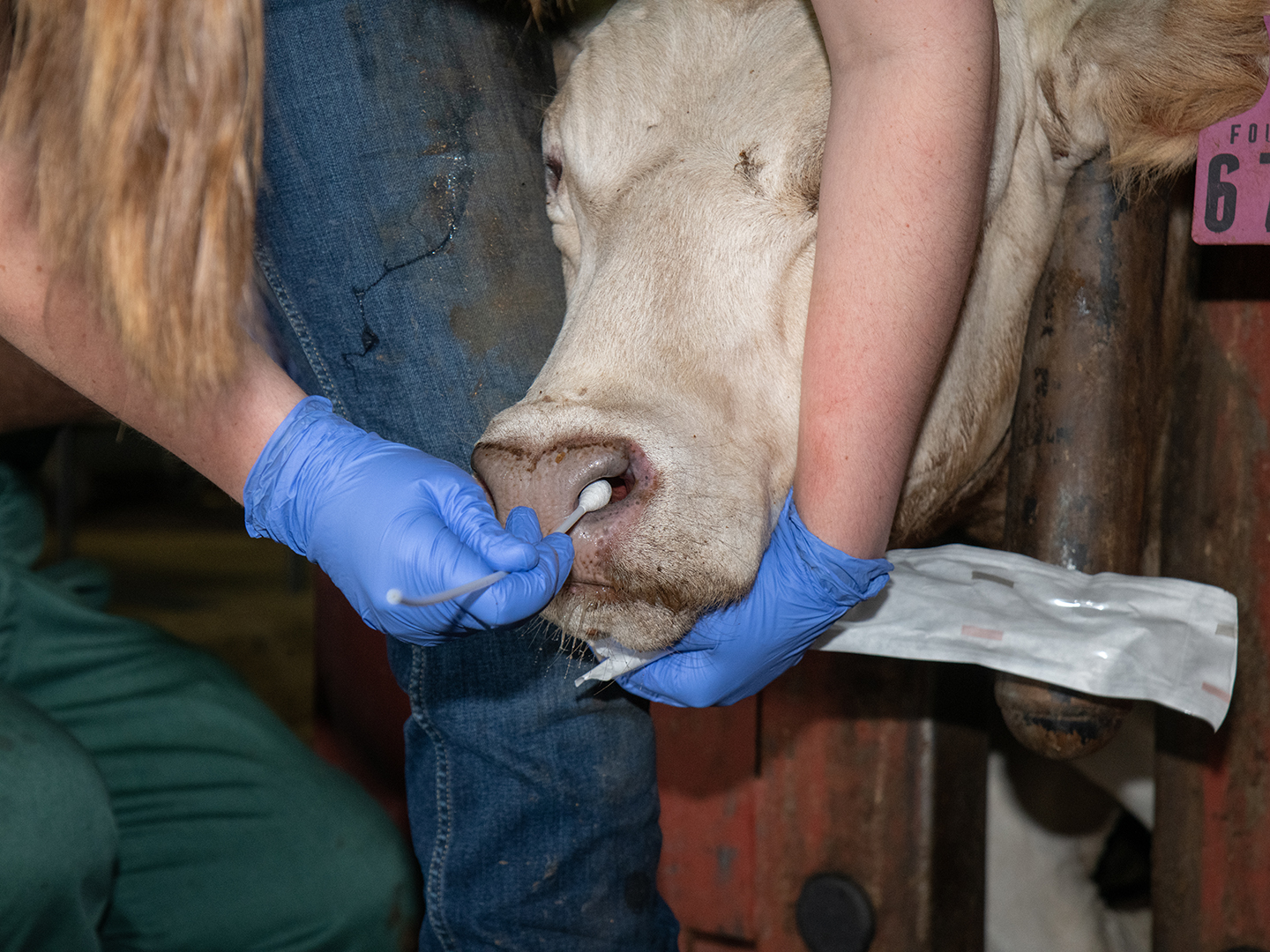


Valeris-Chacin’s goal with this grant is to create a diagnostic tool that can identify the different strains of M. bovis present in a sample; to solve the problem of the scarcity of the bacteria, he will be modifying a process called target enrichment that has worked for other pathogens.
“Using this technique, we can concentrate the DNA of the M. bovis and then hopefully that will give us enough information to do strain-level detection,” he said. “Doing that would open the door for a lot of information that we just don’t have right now. So we’re trying to develop a new technique that will give us that information.”
Being able to identify specific strains, and, therefore, better understand their roles in the different stages of BRD, would be an important step toward a long-term goal of this field, the ability to better track, and ultimately predict, the spread of BRD.
“In the future we might be able to go into a feedlot that’s early on in a BRD issue, sample those animals, detect problematic strains, and begin treatment to minimize the effects before most of the animals are even showing symptoms,” he said.
More precise treatment of BRD ties into another major focus area of VERO research—better use of antibiotics. There is ongoing research into whether antibiotic use in livestock affects humans, but beyond that, treating herds of thousands of animals is expensive and BRD treatment is one of the leading causes of antibiotic use in cattle.
“There is definitely a lot of interest in being able to use antibiotics to prevent or treat BRD in only the animals that will benefit the most,” he said. “But to do that, we’re going to need a lot of information we don’t have right now.”
Though this project focuses on a very specific aspect of BRD, Valeris-Chacin hopes that the groundwork he lays with M. bovis can be translated to other pathogens and ultimately lead to new approaches in BRD research.
“We need to keep studying these bacteria behind BRD because the knowledge we have is incomplete,” he said. “The greatest proof of that is the number of animals that get sick hasn’t declined. We have treatments and some prevention measures, but they’re insufficient. This strain-level data is missing in all of the BRD pathogens, but particularly in M. bovis.”
###
For more information about the Texas A&M School of Veterinary Medicine & Biomedical Sciences, please visit our website at vetmed.tamu.edu or join us on Facebook, Instagram, and Twitter.
Contact Information: Jennifer Gauntt, Director of VMBS Communications, Texas A&M School of Veterinary Medicine & Biomedical Sciences, jgauntt@cvm.tamu.edu, 979-862-4216


